AbstractBackground/AimsEndoscopic ultrasound-guided tissue acquisition (EUS-TA) is a standard diagnostic method for biliary tract cancer (BTC), and samples obtained in this manner may be used for comprehensive genomic profiling (CGP). This study evaluated the utility of EUS-TA for CGP in a clinical setting and determined the factors associated with the adequacy of CGP in patients with BTC.
MethodsCGP was attempted for 105 samples from 94 patients with BTC at the Aichi Cancer Center, Japan, from October 2019 to April 2022.
ResultsOverall, 77.1% (81/105) of the samples were adequate for CGP. For 22-G or 19-G fine-needle biopsy (FNB), the sample adequacy was 85.7% (36/42), which was similar to that of surgical specimens (94%, p=0.45). Univariate analysis revealed that 22-G or larger FNB needle usage (86%, p=0.003), the target primary lesions (88%, p=0.015), a target size Ōēź30 mm (100%, p=0.0013), and number of punctures (90%, p=0.016) were significantly positively associated with CGP sample adequacy.
INTRODUCTIONBiliary tract cancer (BTC) is the sixth leading cause of cancer mortality and approximately 17,000 people with BTC die annually in Japan. According to data from 2009 to 2011 in Japan, the five-year survival rate is 24.5%.1
Although surgical resection is the only curative therapy, half of the BTC cases are unresectable at the time of diagnosis. According to the results from the Japanese Biliary Tract Cancer Statistics Registry from 1998 to 2004, the curative resection rate was 47.3% (752/1,590) in patients with gallbladder cancer (GBC) and 46.7% (884/1,894) in those with bile duct cancer.2 Hence, chemotherapy is the standard therapy for BTCs.
The first-line chemotherapy is a cisplatin-gemcitabine-based treatment, yielding a median overall survival period of 11.7 months in a cisplatin-gemcitabine group (ABC-02 trial),3 13.5 months in a cisplatin-gemcitabine plus tegafur/gimeracil/oteracil group (KHBO1401-MITSUBA trial),4 and 16.8 months in a cisplatin-gemcitabine plus durvalumab group.5
Combination chemotherapy of 5-fluorouracil and oxaliplatin,6 and liposomal irinotecan in combination with fluorouracil and leucovorin,7 have been recommended in the National Comprehensive Cancer Network (NCCN) guidelines as second-line chemotherapy after the failure of cisplatin-gemcitabine chemotherapy. However, as no evidence-based second-line chemotherapy is currently recommended for BTC, the gemcitabine plus cisplatin-based treatment is, in effect, also its last-line treatment. The small array of chemotherapies available for BTC contributes to its poor prognosis; therefore, alternative chemotherapeutic drugs are urgently required.
The NCCN guidelines recommend that molecular analysis be performed when a patient is first diagnosed with unresectable BTC. Comprehensive genomic profiling (CGP) has been approved in Japan since June 2019, and nearly 38.9% (93/239) of Japanese patients with BTCs in one study harbored genetic variants that were potential therapeutic targets.8 Certain targeted molecular therapies, such as pemigatinib for fibroblast growth factor receptor 2 (FGFR2) fusions and ivosidenib for isocitrate dehydrogenase-1 (IDH1) mutations, have been granted accelerated approval by the United States Food and Drug Administration, and pemigatinib has been approved in Japan since June 2021. Thus, precision medicine has been spreading not only in medical research but also in clinical practice.
Endoscopic ultrasound-guided tissue acquisition (EUS-TA) has been useful for BTC, and a previous meta-analysis involving 957 patients with BTC demonstrated that the sensitivity and specificity of EUS-TA were 80% (95% confidence interval [CI], 74%ŌĆō86%) and 97% (95% CI, 94%ŌĆō99%), respectively.9 EUS-TA has already been used to obtain tissue samples for CGP, for example, in patients with pancreatic cancers. However, to the best of our knowledge, there are no reports describing the best method for obtaining CGP samples from patients with BTC. Therefore, we conducted this study with the aim of evaluating the utility of EUS-TA for CGP in a clinical setting and determining the factors associated with the adequacy of CGP in patients with BTC.
METHODSPopulationTissue samples were obtained for tissue-based CGP analysis at the Aichi Cancer Center Hospital, Japan, from October 2019 to December 2022. Demographic and clinical information of all patients were retrospectively collected from the Aichi Cancer Center Hospital Clinical Database.
Endoscopic procedureAll EUS-TA procedures were performed using linear array endoscopes (Olympus GF-UCT260, Olympus Medical Systems or EG-580UT or GF-740UT, Fujifilm). We used EZ Shot 3 Plus needles (Olympus Medical Systems) for fine-needle aspiration (FNA) and Acquire needles (Boston Scientific) for fine-needle biopsy (FNB). We routinely performed three punctures per patient and used rapid on-site evaluation to verify the viability of the tumor tissue and prevent contamination with interstitial, necrotic, or connective tissues. However, we adjusted the number of times by visually examining the white specimens of the samples taken. Each tissue sample was stored in a 10% formalin solution.
Pathological evaluationAt the Aichi Cancer Center Hospital, pathologists evaluated the tumor volume and cellularity of each sample by microscopic examination to determine their appropriateness for CGP analysis. We performed three genomic panel tests using the following: FoundationOne CDx (Foundation Medicine), OncoGuide NCC Oncopanel System (Sysmex Corporation), and Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT; Memorial Sloan Kettering Cancer Center).
1) Test criteria(1) Foundation One CDxIf the total tumor surface area was Ōēź25 mm2 (5├Ś5 mm), it was divided into (1) one formalin-fixed paraffin-embedded block and one hematoxylin and eosin (H&E)-stained slide or (2) 10 unstained slides (positively-charged and unbaked, 4ŌĆō5 ┬Ąm thick) and one H&E-stained slide. If the tumor surface area was <25 mm2, it was divided into 10 unstained slides (4ŌĆō5 ┬Ąm thick) to achieve a tissue volume of 1 mm3. Samples were only used if they had Ōēź20% tumor nuclei content (total number of tumor cells divided by number of cells with nuclei), although Ōēź30% is optimal.
(2) OncoGuide NCC Oncopanel SystemSamples had to meet three criteria: (1) five unstained slides (10 ┬Ąm thick) could be made; (2) total tumor surface area was Ōēź16 mm2 (4├Ś4 mm), and (3) percent tumor nuclei content was Ōēź20%.
(3) MSK-IMPACTSamples had to meet three criteria: (1) 20 unstained slides (4ŌĆō5 ┬Ąm) could be made; (2) total tumor surface area was Ōēź25 mm2 (5├Ś5 mm), and (3) percent tumor nuclei content was Ōēź10%.
In this study, samples with <20% tumor nuclei content, insufficient material, or unanalyzable DNA were considered inadequate. Samples for which CGP was successfully performed were considered adequate.
If the tissue criteria were met, MSK-IMPACT panel tests were performed according to the patientŌĆÖs wishes because MSK-IMPACT panel tests are not covered by insurance in Japan.
The Foundation One CDx panel test can detect a large number of genetic alterations compared to the OncoGuide NCC Oncopanel tests. If the samples met the relevant criteria, Foundation One CDx panel tests were performed.
Statistical analysesWe used the chi-square test or FisherŌĆÖs exact test to compare categorical variables, with two-sided p-values <0.05 considered as statistically significantly. The area under the receiver operating characteristic (ROC) curve (AUC) was calculated to determine the optimal target size for CGP. All statistical analyses were performed using Microsoft Excel 2017 (Microsoft Corporation).
EndpointsThe primary endpoint was sample adequacy for CGP. The secondary endpoints were the results of the CGP analysis and the occurrence of adverse events, which were graded according to the lexicon of the American Society for Gastrointestinal Endoscopy.10
RESULTSPatient characteristicsWe collected 137 samples from 126 patients for prescreening (Fig. 1). The pathologists judged 113 samples as meeting the suitability criteria and 24 as inadequate. We further excluded 32 samples for which tissue-based CGP was not performed, even though they met the criteria. The remaining 81 samples (81 patients) were considered adequate. We investigated 81 adequate and 24 inadequate samples obtained from 94 patients, the characteristics of whom are summarized in Table 1. The median age of the patients was 65 years, and 46 of 94 patients were women. The primary tumors were intrahepatic cholangiocarcinoma (ICC) (n=37), extrahepatic cholangiocarcinoma (ECC) (n=17), GBC (n=34), gallbladder neuroendocrine carcinoma (n=2), papillary neuroendocrine carcinoma (n=2), papillary adenocarcinoma (n=1), and bile duct neuroendocrine tumor (n=1).
In this study, we obtained tissue samples from six patients with ECC using EUS-TA, and punctured the thick bile duct wall (n=4) and the liver invasion area (n=2).
CGP was performed most frequently using the Foundation One CDx panel (n=41), followed by the OncoGuide NCC Oncopanel System (n=36) and the MSK-IMPACT panel (n=4).
In seven cases, prescreening was performed twice or thrice, and 18 samples were obtained from these seven patients (Table 1).
Sample adequacy
Table 2 summarizes the sample adequacy for CGP for each sampling method. Overall, 77.1% (81/105) of the samples were adequate for CGP. The sample adequacy of the surgical specimens was 93.8% (30/32), and that of the specimens acquired by EUS-TA and percutaneous liver biopsy were 76.5% (39/51) and 64.7% (11/17), respectively.
Table 3 summarizes the sample adequacy of specimens obtained via EUS-TA, with comparisons based on various factors, including gauge (19-G, 22-G, and 25-G), needle type (FNB or FNA), targets (liver lesions or not, primary or metastasis), target size (Ōēź30 mm or <30 mm), number of punctures (Ōēź3 or <3 times), and puncture route (transgastric or transduodenal). Accordingly, needle gauge (19-G vs. 22-G vs. 25-G: 93% vs. 60% vs. 0%, p=0.013) and type (FNB vs. FNA: 84% vs. 38%, p=0.013) were associated with sample adequacy. The adequacy rate of samples obtained using 19-G or 22-G (Ōēź22-G) FNB needles was 85.7% (36/42), which was higher than that of the samples obtained using other needles (33%, p<0.001). The adequacy rate did not differ between samples obtained with Ōēź22-G FNB needles and surgical specimens (86% vs. 94%, p=0.45).
Other univariate analyses revealed that the primary lesion (primary vs. metastasis: 88% vs. 56%, p=0.015), target size (Ōēź30 mm vs. <30 mm: 100% vs. 43%, p<0.001), and number of punctures (Ōēź3 times vs. <3 times: 90% vs. 57%, p=0.016) were associated with sample adequacy, whereas liver lesions (liver vs. others: 84% vs. 63%, p=0.10) and puncture route (transgastric vs. transduodenal: 68% vs. 83%, p=0.32) were not. We analyzed the ROC curve and defined 30 mm as the cut-off index for the target size (AUC=0.86, p<0.001). The largest target size was 95 mm, and the lesion was the liver invasion area of the GBC.
A multivariate analysis was not performed because the number of samples obtained using EUS-TA was limited.
In the group in whom 19-G FNB needles were used, 44.8% (13/29) of the samples met the suitability criteria for the Foundation One CDx panel and 6.9% (2/29) met the criteria for the MSK-IMPACT panel. 19-G FNB was available to perform Foundation One CDx more than the other needles (19-G FNB vs. 22-G FNB vs. others: 45% vs. 15% vs. 0%, p=0.016). Genomic panel tests requiring a larger amount of tumor tissue, such as the Foundation One CDx or MSK-IMPACT panels, were more challenging to perform (Table 4).
Complications of EUS-TAThree patients (6%) experienced complications: one had mild abdominal pain (19-G FNB, ICC, liver lesion), one had mild cholangitis (19-G FNB, GBC, gallbladder wall), and one had mild pancreatitis (22-G FNA, ICC, liver lesion). No moderate, severe, or fatal complications were observed. No differences in the complication rates were observed according to needle gauge (19-G vs. 22-G vs. 25-G: 7% vs. 5% vs. 0%, p>0.99) or type (FNA vs. FNB: 13% vs. 5%, p=0.41).
CGP resultsThe CGP results are illustrated in Figure 2. Based on the NCCN guidelines, we identified eight therapeutic molecular markers: IDH1 mutations, FGFR2 fusions, neurotrophic receptor tyrosine kinase fusions, BRAF V600E mutation, receptor tyrosine-protein kinase erbB-2 (ERBB2) amplification, rearrangement during transfection fusions, microsatellite instability-high status, and tumor mutational burden-high status.
The incidence of IDH1 mutations was 9.7% (3/31) among patients with ICC and 0% in patients with ECC or GBC. FGFR2 fusions were detected only in patients with ICC, at a rate of 12.9% (4/31). ERBB2 amplification was detected in 21.4% (6/28) of patients with GBC. Any of the aforementioned targeted therapeutic molecular markers were detected in 22.6% (7/31) of patients with ICC and in 32.1% (9/28) of those with GBC, but in none of the patients with ECC (Table 5).
Kirsten rat sarcoma virus (KRAS) mutations were detected in 32.3% (10/31), 35.7% (5/14), and 7.1% (2/28) of patients with ICC, ECC, and GBC, respectively. Patients with GBC had a lower incidence of KRAS mutations than those with cholangiocarcinoma (7% vs. 33%, p=0.011). Finally, TP53 mutations were detected in 32.3% (10/31), 71.4% (10/14), and 53.6% (15/28) of patients with ICC, ECC, and GBC, respectively.
DISCUSSIONEUS-TA is a useful diagnostic method for BTC. Since most BTCs are unresectable at diagnosis, tissue samples must be obtained in an alternative manner. Percutaneous liver biopsy is widely used for diagnosis and to obtain samples. However, in this study, only 65% of the samples obtained in this manner were adequate for CGP analysis. However, 86% of the samples obtained using 22-G or 19-G FNB needles were adequate for CGP. EUS-TA enables the collection of samples not only from liver lesions, but also from lymph nodes and gallbladder walls more easily than percutaneous biopsy.
Transpapillary tissue sampling is a good method for diagnosis, but its sensitivity is 48.1%11 and such samples may involve normal biliary duct epithelium. This may reduce the tumor cellularity of the samples, therefore, transpapillary samples were not used for tissue-based CGP.
Since the genetic analysis of samples obtained using EUS-TA was first reported in 2001,12 its use has become widespread. The pancreas (65%) and lungs (26%) are the most common sites of primary tumors among patients undergoing EUS-TA.13 However, very few reports have been published on the genetic analysis of BTC samples obtained using EUS-TA.
The first report of genetic analysis of BTC using specimens obtained through EUS-FNA was published in 2019. Therein, Hirata et al.14 reported that 95.2% (20/21) of EUS-FNA samples were successfully analyzed using 22-G (n=19) or 25-G needles (n=2) (Expect or Acquire, Boston Scientific). The Iron AmpliSeq Cancer Hotspot Panel v2 (Thermo Fisher Scientific) was used for genomic analysis.
To the best of our knowledge, ours is the first report on the genetic analysis of EUS-TA samples using genomic panel tests, such as the Foundation One CDx and OncoGuide NCC Oncopanel System, in a clinical setting. In this study, 76.5% (39/51) of the EUS-TA samples and 85.7% (36/42) of those obtained using 22-G or 19-G FNB needles were adequate for CGP analysis. Moreover, samples obtained using 22-G or 19-G FNB yielded a high adequacy rate, similar to that of surgical specimens.
Other primary tumors, including pancreatic cancers, may also be sampled for CGP. Ikeda et al.15 revealed that 19-G needles (19-G vs. 22-G; 56% vs. 23%, p<0.001) and FNB needles (FNB vs. FNA: 48% vs. 11%, p<0.001) were significantly and positively associated with the adequacy of the OncoGuide NCC Oncopanel System. In their study, the multivariate analysis revealed that the independent factors associated with adequacy were needle gauge (19-G; odds ratio [OR], 2.53; 95% CI, 1.15ŌĆō5.57; p=0.021) and needle type (FNB; OR, 3.57; 95% CI, 1.05ŌĆō12.20; p=0.041).
Takahashi et al.16 also reported that the amount of tissue obtained by 19-G FNB was approximately three times larger than that obtained by 19-G FNA (median 15.20 mm2 vs. 5.44 mm2, p=0.010) and that obtained by 22-G FNB (median 15.20 mm2 vs. 4.49 mm2, p=0.008). Both studies indicated that 19-G and FNB needles were preferred over other needles for patients with pancreatic cancer. Our results were consistent with previous results in that large-gauge and FNB needles had a significant association with adequacy (19-G vs. 22-G; 93% vs. 55%, p=0.0021; and FNB vs. FNA:84% vs. 33%, p=0.013).
In the present study, IDH1 mutations and FGFR2 fusions were detected in patients with ICC, and ERBB2 amplifications were detected in patients with GBC. According to previous reports, the prevalence of IDH1 mutations and FGFR2 fusions is 13.1%17 and 9% to 14%, respectively, among patients with ICC,18 and that of ERBB2 amplifications is 12.8% among patients with GBC.19 These three targetable alterations were the most frequently detected alterations among patients with BTC in the current study. Israel et al.20 demonstrated that, among patients with ICC, FGFR2 fusions were detected less frequently in liquid biopsies than in tumor tissues.
Berchuck et al.21 reported on the clinical landscape of cell-free DNA (cfDNA) alterations in patients with advanced BTC. The landscape of cfDNA (n=1671) and tissue (n=349, AACR Project GENIE database of metastatic BTC) differed significantly in terms of the detection of FGFR2 fusions. No significant differences in the frequencies of IDH1 mutations or ERBB2 amplifications were observed between cfDNA and tissue samples in this study. Therefore, we believe that tissue-based CGP is preferable to liquid-based CGP for detecting FGFR2 fusions. Furthermore, pemigatinib has been approved in Japan since 2022, and tissue-based CGP is preferred especially for patients with ICC.
This study had a some limitations. First, this was a retrospective, single-center study. Second, there was a risk of selection bias. Third, there might have been some cases in which tissue-based CGP could be performed; however, was not performed CGP because the pathologists judged the samples to be inadequate for analysis. In the future, we need to perform a multicenter study with a large cohort of patients with BTC to investigate the efficacy of tissue-based CGP using samples obtained by EUS-TA and to identify the factors associated with sample adequacy.
In conclusion, EUS-TA is safe for obtaining tissue samples for CGP. Factors significantly positively associated with adequacy were the use of 22-G or 19-G FNB needles, a target size Ōēź30 mm, the target of primary lesions, and the number of punctures. Patients with BTC frequently exhibit druggable genetic alterations, with FGFR2 fusions detected more frequently than other alterations in tissue samples. We believe that EUS-TA is useful for tissue-based CGP of BTC, particularly in patients with ICC.
NOTESConflicts of Interest
Dr. Mizuno reports grants and fees from Yakult Honsha, Novartis, MSD, Ono Pharmaceutical, ASLAN Pharmaceuticals, Incyte, Seagen, Taiho Pharmaceutical and Dainippon Sumitomo Pharma.; none of the grants are connected with the submitted work. The authors have no potential conflicts of interest.
Fig.┬Ā2.Genomic results of tissue-based CGP. CGP, comprehensive genomic profiling; ICC, intrahepatic cholangiocarcinoma; ECC, extrahepatic cholangiocarcinoma; GBC, gallbladder cancer; GN, gallbladder neuroendocrine carcinoma; PN, papilla neuroendocrine carcinoma; PA, papilla adenocarcinoma; BN, bile duct neuroendocrine tumor; IDH, isocitrate dehydrogenase; FGFR, fibroblast growth factor receptor; PIK3CA, phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; ERBB, receptor tyrosine-protein kinase erbB; BRCA, breast cancer susceptibility gene; MDM, murine double minute; CDKN, cyclin dependent kinase inhibitor; CDK, cyclin dependent kinase; NTKR, neurotrophic receptor tyrosine kinase; ALK, anaplastic lymphoma kinase; MSH, mutS homolog; KRAS, Kirsten rat sarcoma virus; ARID, AT-rich interactive domain-containing protein; MSI-H, microsatellite instability-high; TMB-H, tumor mutational burden-high. 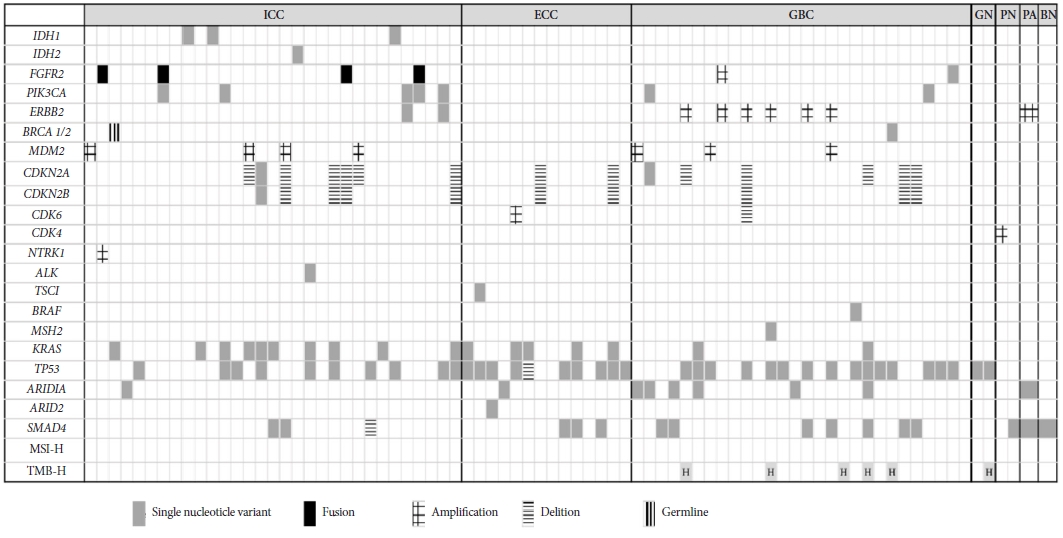
Table┬Ā1.Patient characteristics Values are presented as median (range) or number only. EUS-TA, endoscopic ultrasound-guided tissue acquisition; MSK-IMPACT, Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets. a)Other samples: skin biopsy (2), bronchoscopic biopsy (1), endoscopic biopsy (1), computed tomography guided biopsy (1). Table┬Ā2.Sample adequacy Table┬Ā3.Factors associated with samples adequacy Table┬Ā4.EUS-TA and genomic panel tests Table┬Ā5.Therapeutic molecular makers Values are presented as number (%). ICC, intrahepatic cholangiocarcinoma; GBC, gallbladder cancer; ECC, extrahepatic cholangiocarcinoma; IDH1, isocitrate dehydrogenase-1; FGFR2, fibroblast growth factor receptor 2; NTRK, neurotrophic receptor tyrosine kinase; ERBB2, receptor tyrosine-protein kinase erbB 2; RET, rearranged during transfection; MSI-H, microsatellite instability-high; TMB-high, tumor mutational burden-high. REFERENCES1. Foundation for Promotion of Cancer Research. Cancer Statistics in Japan 2022. National Cancer Center; 2022;[cited 2022 Dec 28]. Available from: https://ganjoho.jp/public/qa_links/report/statistics/2022_en.html.
2. Miyakawa S, Ishihara S, Horiguchi A, et al. Biliary tract cancer treatment: 5,584 results from the Biliary Tract Cancer Statistics Registry from 1998 to 2004 in Japan. J Hepatobiliary Pancreat Surg 2009;16:1ŌĆō7.
3. Valle J, Wasan H, Palmer DH, et al. Cisplatin plus gemcitabine versus gemcitabine for biliary tract cancer. N Engl J Med 2010;362:1273ŌĆō1281.
4. Ioka T, Kanai M, Kobayashi S, et al. Randomized phase III study of gemcitabine, cisplatin plus S-1 versus gemcitabine, cisplatin for advanced biliary tract cancer (KHBO1401-MITSUBA). J Hepatobiliary Pancreat Sci 2023;30:102ŌĆō110.
5. Oh DY, He AR, Qin S, et al. A phase 3 randomized, double-blind, placebo-controlled study of durvalumab in combination with gemcitabine plus cisplatin (GemCis) in patients (pts) with advanced biliary tract cancer (BTC): TOPAZ-1. J Clin Oncol. 2022;40(4_suppl):378.
6. Lamarca A, Palmer DH, Wasan HS, et al. Second-line FOLFOX chemotherapy versus active symptom control for advanced biliary tract cancer (ABC-06): a phase 3, open-label, randomised, controlled trial. Lancet Oncol 2021;22:690ŌĆō701.
7. Yoo C, Kim KP, Jeong JH, et al. Liposomal irinotecan plus fluorouracil and leucovorin versus fluorouracil and leucovorin for metastatic biliary tract cancer after progression on gemcitabine plus cisplatin (NIFTY): a multicentre, open-label, randomised, phase 2b study. Lancet Oncol 2021;22:1560ŌĆō1572.
8. Nakamura H, Arai Y, Totoki Y, et al. Genomic spectra of biliary tract cancer. Nat Genet 2015;47:1003ŌĆō1010.
9. Sadeghi A, Mohamadnejad M, Islami F, et al. Diagnostic yield of EUS-guided FNA for malignant biliary stricture: a systematic review and meta-analysis. Gastrointest Endosc 2016;83:290ŌĆō298.
10. Cotton PB, Eisen GM, Aabakken L, et al. A lexicon for endoscopic adverse events: report of an ASGE workshop. Gastrointest Endosc 2010;71:446ŌĆō454.
11. Navaneethan U, Njei B, Lourdusamy V, et al. Comparative effectiveness of biliary brush cytology and intraductal biopsy for detection of malignant biliary strictures: a systematic review and meta-analysis. Gastrointest Endosc 2015;81:168ŌĆō176.
12. Rader AE, Avery A, Wait CL, et al. Fine-needle aspiration biopsy diagnosis of gastrointestinal stromal tumors using morphology, immunocytochemistry, and mutational analysis of c-kit. Cancer 2001;93:269ŌĆō275.
13. Kuwatani M, Sakamoto N. Evolution and a promising role of EUS-FNA in gene and future analyses. Endosc Ultrasound 2020;9:151ŌĆō153.
14. Hirata K, Kuwatani M, Suda G, et al. A novel approach for the genetic analysis of biliary tract cancer specimens obtained through endoscopic ultrasound-guided fine needle aspiration using targeted amplicon sequencing. Clin Transl Gastroenterol 2019;10:e00022.
15. Ikeda G, Hijioka S, Nagashio Y, et al. Fine-needle biopsy with 19G needle is effective in combination with endoscopic ultrasound-guided tissue acquisition for genomic profiling of unresectable pancreatic cancer. Dig Endosc 2023;35:124ŌĆō133.
16. Takahashi K, Yasuda I, Hanaoka T, et al. Comparison of histological sample volumes among various endoscopic ultrasound-guided biopsy needles. J Clin Med 2021;10:3560.
17. Boscoe AN, Rolland C, Kelley RK. Frequency and prognostic significance of isocitrate dehydrogenase 1 mutations in cholangiocarcinoma: a systematic literature review. J Gastrointest Oncol 2019;10:751ŌĆō765.
18. Maruki Y, Morizane C, Arai Y, et al. Molecular detection and clinicopathological characteristics of advanced/recurrent biliary tract carcinomas harboring the FGFR2 rearrangements: a prospective observational study (PRELUDE Study). J Gastroenterol 2021;56:250ŌĆō260.
19. Roa I, de Toro G, Schalper K, et al. Overexpression of the HER2/neu gene: a new therapeutic possibility for patients with advanced gallbladder cancer. Gastrointest Cancer Res 2014;7:42ŌĆō48.
|
|
|||||||||||||||||||||||||||||||||||||
